
Gene silencing refers to a collection of mechanisms that cells use to turn off or suppress the expression of specific genes. By preventing the transcription or translation of a gene, gene silencing provides an important method of controlling gene expression and regulating cellular processes.
Understanding gene silencing mechanisms has revolutionized molecular biology research and provided the basis for novel therapeutic approaches.
Definition and Importance of Gene Silencing
Gene silencing describes the process by which cells prevent the expression of a particular gene. It acts as an “off switch” that inhibits transcription or translation of a specific gene sequence. This silencing can occur through epigenetic modifications to DNA or histones that prevent transcription or by mRNA degradation or interference that blocks translation.
By selectively silencing genes, cells exert dynamic control over gene expression, only synthesizing proteins when and where they are needed. This regulation is crucial for normal growth, development, and health. Defects in silencing mechanisms are linked to cancer, autoimmune disorders, and metabolic diseases.
Researchers rely on artificially inducing gene silencing to study gene function. Clinicians are exploring gene silencing techniques as a basis for novel gene therapies that can treat medical conditions by silencing disease-causing genes. The discovery of gene silencing has revolutionized molecular genetics and provided insight into fundamental cellular processes.
Historical Context of Gene Silencing Research
Richard Jorgensen conducted the foundational research into gene silencing in the late 1980s. Studying petunias, he unexpectedly found that introducing additional copies of a pigmentation gene led to silencing both endogenous and introduced genes rather than increasing flower color. This phenomenon was termed “co-suppression”.
In the 1990s, Andrew Fire and Craig Mello identified double-stranded RNA as the trigger for RNA interference (RNAi) and sequence-specific post-transcriptional gene silencing in worms. Around the same time, plant biologists discovered small interfering RNAs (siRNAs) that directed mRNA degradation.
These crucial discoveries marked the emergence of RNAi as a mechanism of gene silencing guided by small non-coding RNAs. In 1998, Andrew Fire and Craig Mello received the Nobel Prize for their seminal research elucidating the process of RNAi in C. elegans.
In addition to RNAi, other key advances in the 1990s included identifying miRNA regulation and DNA methylation’s role in gene silencing. The past three decades of intensive research have uncovered the diverse molecular processes that allow cells to silence genes at the transcriptional and post-transcriptional levels selectively.
Mechanisms of Gene Silencing
There are two major types of gene silencing:
Transcriptional Gene Silencing
This silencing acts at the DNA level to prevent gene transcription into mRNA. The two main methods cells use are:
A. DNA Methylation
DNA methylation involves adding a methyl group to cytosine bases, often in areas called CpG islands in gene promoters. DNA methyltransferase (DNMT) enzymes catalyze this modification. Methylation typically leads to gene inactivation by blocking the binding of transcription factors.
B. Histone Modification
Histone proteins help package DNA into chromatin. Modifications like deacetylation and methylation of histone tails alter chromatin structure, leading to tighter packing that reduces gene accessibility and transcription.
Post-Transcriptional Gene Silencing
The Mechanism of PTGS processes is referred to, which takes place only after the transcription of the genetic information from the DNA to the transmitting mRNA, before and during the translation. Among the forms of PTGS include in particular the nonsense-mediated decay (NMD) and RNA interference (RNAi).
During the nonsense-mediated mRNA, priority decay avoiding nonsense point mutations used is the RNA interference a predominantly regulatory process with the participation of specific RNA molecules such as miRNA and siRNA. The PTGS may lead to an intensified degradation of the mRNA of a particular gene.
Due to the degradation of the mRNA translation, and thus the formation of the species is the gene product (usually a protein) prevents. Moreover, a gene-specific direct inhibition of translation as a result of post-transcriptional gene silencing is possible.
This type of silencing blocks protein production from an mRNA transcript. Major mechanisms include:
A. MicroRNA (miRNA) Silencing
miRNAs are short, non-coding RNAs that bind to complementary mRNA sequences. It leads to mRNA degradation or translation blocking, suppressing protein output.
B. Small Interfering RNA (siRNA) Silencing
siRNAs have a similar function as miRNAs, directing cleavage of target mRNAs in a sequence-specific manner through RNAi. siRNAs are typically exogenous or artificially introduced.
By employing these various techniques, cells can fine-tune gene expression in response to biological cues and environmental conditions. Next, we’ll explore the mechanisms underlying transcriptional and post-transcriptional gene silencing in more detail.
Transcriptional Gene Silencing
Transcriptional gene silencing acts directly on a gene’s DNA sequence to prevent its transcription into mRNA and protein. The two main methods cells use are DNA methylation and histone modification.
A transcriptional gene silencing is the result of epigenetic changes in DNA (like DNA methylation or histone modifications) or by the binding of repressors to a Silencer or by non-coding RNA. Due to the modifications of Histone and is a kind of heterochromatic state created by the gene which it transcription machine (RNA polymerase, transcription factors to bind, etc.) closed.
The classic example is the as position effect variegation (PEV) Marked phenomenon. It changes the chromatin state and thereby controls the transcription activity of the gene or the gene region concerned.
DNA Methylation
DNA methylation involves adding a methyl group (-CH3) to cytosine bases in the DNA sequence, converting cytosine to 5-methylcytosine. This modification is catalyzed by enzymes called DNA methyltransferases (DNMTs).
In mammals, DNA methylation primarily occurs at CpG sites, where a cytosine nucleotide is next to a guanine nucleotide in the linear sequence of bases. These CpG sites tend to cluster in CpG islands, often found in gene promoter regions.
DNA Methyltransferases
Three main DNMT enzymes establish and maintain DNA methylation patterns:
- DNMT1 – Copies existing methylation patterns during DNA replication
- DNMT3A and 3B – Add methyl groups to previously unmethylated CpGs
DNMT activity leads to hypermethylation of CpG islands in gene promoters, associated with stable gene silencing. The methyl groups physically impede the binding of transcription factors to gene promoters, blocking transcription initiation.
Role of DNA Methylation in Gene Inactivation
DNA methylation is crucial in regulating gene expression during development and cellular differentiation. Extensive methylation of CpG islands leads to long-term silencing of tissue-specific genes, locking cells into specialized fates.
Abnormal hypermethylation and silencing of tumor suppressor genes is an important step in cancer development. DNA methylation marks provide stable, heritable silencing signals passed on through cell divisions.
Histone Modification
In addition to DNA methylation, gene silencing can occur through histone proteins’ modifications. Histones help package DNA into structural units called nucleosomes that make up chromatin fibers.
Modifications like acetylation, methylation, and phosphorylation of histone tails change the structure and packing of chromatin, altering gene accessibility.
Histone Acetylation and Deacetylation
Histone acetyltransferases (HATs) add acetyl groups to lysine residues on histone tails, neutralizing their positive charge. It loosens chromatin structure, allowing increased transcription.
Histone deacetylases (HDACs) remove those acetyl groups, increasing the positive charge on histones and promoting tighter chromatin packing and reduced transcription.
Histone Methylation
Histone methyltransferases (HMTs) add one or more methyl groups to histone tails, signaling either activation or repression of genes depending on the site and degree of methylation. For example, H3K9 methylation is associated with the formation of heterochromatin and gene silencing.
Role of Histone Modification in Gene Silencing
The combination of histone modifications alters chromatin density, with tight chromatin blocking access for transcription machinery. Histone modification provides flexible, reversible gene silencing compared to more permanent DNA methylation marks.
Post-Transcriptional Gene Silencing
In addition to directly blocking transcription, cells can silence genes after transcription by targeting the mRNA transcripts for destruction or blocking their translation. The main mechanisms for post-transcriptional gene silencing are miRNA regulation and RNA interference directed by siRNAs.
MicroRNA (miRNA) Silencing
MicroRNAs (miRNAs) are short, non-coding RNA molecules, usually about 22 nucleotides long, that bind to complementary sequences in target mRNAs. This binding leads to degradation or translation inhibition of the target mRNA, effectively silencing its output.
Biogenesis of miRNAs
miRNAs are first transcribed as long primary transcripts called pri-miRNAs that fold into hairpin structures. The RNase Drosha processes pri-miRNAs into shorter pre-miRNAs. After export to the cytoplasm, Dicer cleaves the pre-miRNA into a miRNA duplex. One strand is incorporated into the RNA-induced silencing complex (RISC).
Mechanism of miRNA Action
The miRNA guides the RISC complex to bind to partially complementary sites in target mRNAs’ 3′ untranslated region (3′ UTR). Perfect complementarity leads to mRNA cleavage and degradation. Imperfect matches block translation initiation and elongation. A single miRNA can regulate hundreds of target mRNAs.
Target Recognition and Binding Specificity
miRNA target recognition depends on complementarity between the mRNA 3′ UTR sequence and the miRNA seed region (nucleotides 2-8). This imperfect binding allows more promiscuous targeting of multiple mRNAs by each miRNA.
Small Interfering RNA (siRNA) Silencing
Small interfering RNAs (siRNAs) are short double-stranded RNAs that direct cleavage of complementary target mRNAs. They act through a mechanism similar to miRNAs as part of the RNAi pathway. However, siRNAs typically originate from exogenous or introduced dsRNA triggers rather than being encoded endogenously like miRNAs.
Mechanisms and Applications of siRNAs
Inside cells, Dicer processes trigger dsRNAs into siRNA duplexes about 20-25 base pairs long. One strand is incorporated into RISC, which uses the siRNA as a guide to recognize and cleave an mRNA target sequence precisely matching the siRNA. It provides a method for artificially silencing specific genes by introducing complementary siRNAs.
Gene Silencing Techniques
Researchers have developed numerous techniques to induce targeted gene silencing for laboratory studies or potential therapeutic applications. Major approaches include RNA interference, CRISPR systems, and antisense oligonucleotides.
RNA Interference (RNAi)
As described earlier, RNAi refers to any process where introduced double-stranded or small interfering RNAs trigger post-transcriptional silencing of complementary mRNA targets. RNAi techniques include:
- SiRNA transfection – Introducing synthetic siRNA molecules into cells to temporarily knock down target genes.
- ShRNA expression – DNA vectors encode short hairpin RNAs (shRNAs) that get transcribed into siRNA-like molecules inside cells for stable knockdown.
- miRNA overexpression or inhibition – Introducing miRNA mimics or antisense inhibitors.
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)
CRISPR-Cas9 systems provide an efficient way to edit genes by taking advantage of bacterial immune pathways that target and cleave foreign DNA. The Cas9 nuclease is guided by a small guide RNA to induce double-stranded breaks at target sites. When repaired, this damage causes gene knockout. The New versions, like Cas13, can target mRNAs for silencing rather than genomic cleavage.
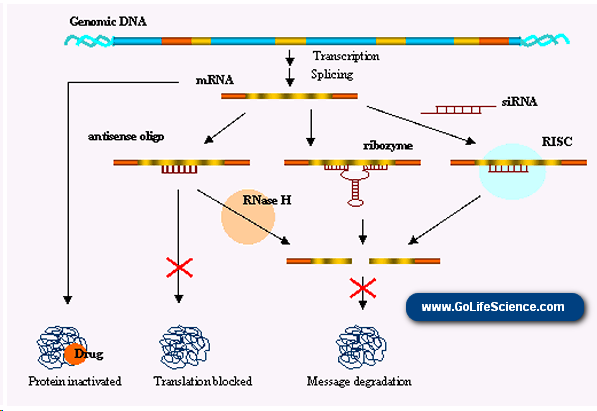
Antisense Oligonucleotides
Antisense oligos are short, synthetic nucleic acids that bind to mRNA transcripts through complementary base pairing. It can induce RNase H cleavage of the mRNA or physically obstruct translation. Modified, enhanced forms using gapmers and LNAs can improve stability and silencing efficiency.
Applications of Gene Silencing
Selective gene silencing through RNAi, CRISPR systems, and antisense oligos provides researchers with powerful tools to control cellular processes and has enabled many invaluable applications.
1. Gene Therapy
Gene silencing techniques can help treat genetic disorders by selectively knocking down the expression of mutated genes. For example, lipid nanoparticle-mediated delivery of siRNAs to liver cells enabled effective silencing of disease-causing genes in patients with transthyretin-mediated amyloidosis.
2. Functional Genomics
Gene silencing via RNAi allows systematic screening for loss-of-function phenotypes. Researchers use siRNA/shRNA libraries to identify genes involved in specific cellular pathways, isolate components affecting drug sensitivity, or determine gene function in normal physiology and disease.
3. Biotechnology and Agriculture
Gene silencing has many uses in biotechnology and agriculture, from enhancing crop traits to producing biologics. For instance, agrochemical companies use RNAi to create pest-resistant plants, while pharmaceutical researchers employ RNAi screens for drug target discovery.
4. Therapeutic Potential in Diseases
In addition to treating genetic disorders, silencing dysregulated genes offers a promising therapeutic avenue for diverse diseases like cancer, viral infections, neurodegenerative conditions, and cardiovascular disorders. Multiple gene-silencing drugs have entered clinical trials. Challenges include improving delivery and avoiding toxicity.
Challenges and Ethical Considerations
While holding great potential, gene silencing techniques also have challenges and ethical concerns that must be carefully considered.
1. Off-Target Effects
One risk is the unintended silencing of non-target genes due to insufficient specificity or the target gene’s sequence homology to other genes. It can cause unexpected side effects and toxicity. Modifications like chemical stabilization help improve siRNA specificity.
2. Ethical Issues in Gene Editing
Powerful gene editing tools raise important ethical issues regarding altering the human genome. Changes made in embryos passed on to future generations are especially controversial. Strict oversight is critical to ensure human gene editing proceeds cautiously and ethically.
3. Regulatory Concerns
Agencies like the FDA and EMA heavily regulate the therapeutic use of gene-silencing technologies. Clinical translation faces hurdles in demonstrating efficacy and safety through extensive testing in animal and human trials.
Future Directions and Research
Researchers continue working to enhance gene silencing techniques and further elucidate cellular mechanisms regulating gene expression. Exciting areas of ongoing research include:
1. Advancements in Gene Silencing Technologies
Novel delivery vehicles, chemical modification strategies, and computational design improvements are expanding the power and specificity of RNAi, antisense oligos, and CRISPR systems for research and therapy.
2. Emerging Trends in Gene Regulation Research
Current frontiers include elucidating new classes of regulatory RNAs like circular RNAs, better-defining chromatin regulation and cooperation between epigenetic controls, and single-cell multi-omics analysis of gene regulatory networks.
Significance of Gene Silencing for Biology and Medicine
Gene silencing research has unlocked new realms of biology by elucidating how cells dynamically regulate genomes through epigenetic modifications and non-coding RNAs. Harnessing natural silencing pathways also enables revolutionary therapeutic approaches to treat diseases by silencing pathological genes. The story of gene silencing illustrates how basic science discoveries transform medicine and fundamentally alter our comprehension of life’s complexities.
Frequently Asked Questions
What is the difference between transcriptional and post-transcriptional gene silencing?
The key difference lies in what stage of gene expression they inhibit. Transcriptional silencing acts directly on the DNA to prevent transcription of the gene sequence into mRNA. It includes DNA methylation and histone modifications that block the gene from being accessed and transcribed. In contrast, post-transcriptional silencing allows transcription but prevents mRNA translation into protein. Mechanisms like RNAi using miRNAs/siRNAs target the mRNA for degradation or obstruct its translation after transcribing it.
How does DNA methylation lead to gene silencing?
DNA methylation involves adding methyl groups to CpG dinucleotides, often clustered in gene promoter regions called CpG islands. This modification leads to gene silencing by a few mechanisms:
– Methyl groups physically impede the binding of transcription factors required to initiate mRNA synthesis.
– Methylated DNA recruits proteins like MeCP2 that further repress transcription through chromatin remodeling.
– Heavily methylated genes accumulate repressive histone marks over time that stably lock in the silent state.
What is the difference between miRNAs and siRNAs?
miRNAs and siRNAs have overlapping functions and mechanisms – both direct post-transcriptional silencing of target genes through RNAi pathways. However, there are a few key differences:
– Origin: miRNAs are produced from endogenous genes; siRNAs come from exogenous dsRNA.
– Complementarity: miRNAs have imperfect binding to targets, while siRNAs match their mRNA targets perfectly.
– Range: Each miRNA regulates multiple genes; siRNAs silence individual targets.
How does CRISPR-Cas9 mediate gene silencing?
The Cas9 nuclease causes double-stranded breaks at targeted gene sequences guided by a small guide RNA. The resulting DNA damage causes gene knockout when repaired via error-prone non-homologous end joining. Newer versions like Cas13 are RNA-guided and can directly cleave mRNAs instead of genomic DNA. Researchers can also engineer “dead” Cas9 lacking nuclease activity to bind genes and block their transcription through steric interference specifically.
What are some challenges facing the therapeutic use of gene silencing?
– Off-target effects leading to toxicity
– Ensuring efficient delivery to target tissues
– Avoiding immune stimulation
– Stability and uptake of RNAs/oligos in circulation
– High cost of synthesis and formulation
– Ethical concerns related to gene editing
– Extensive regulatory testing is required for approval.
Advances in chemistry, delivery materials, and computational modeling continue to help address these hurdles and facilitate the translation of gene-silencing technologies to the clinic. Careful dosing and strict oversight also help ensure their safe and ethical use.
Conclusion
The discovery of diverse gene silencing mechanisms has fundamentally transformed our understanding of gene regulation and provided scientists with versatile new techniques to modify cellular processes. Key points covered here include:
Recap of Key Points
- Gene silencing provides crucial control over gene expression by blocking transcription or translation of specific genes.
- Major mechanisms include DNA methylation, histone modification, RNA interference by miRNAs/siRNAs, and antisense oligonucleotides.
- Researchers can dissect cellular pathways and develop novel therapeutics by selectively inhibiting genes.
- Challenges remain in enhancing specificity and delivery while ensuring ethical applications.
