In molecular biology, an operon is a fundamental unit of gene regulation in prokaryotic cells. Discovered in Escherichia coli by François Jacob and Jacques Monod, the operon model explains how bacteria regulate gene expression efficiently. This process ensures that genes responsible for specific functions, such as metabolism, are activated or deactivated based on environmental conditions, optimizing energy usage.
The concept of operons plays a pivotal role in understanding gene regulation and prokaryotic gene expression. Whether you’re a student diving into the intricacies of bacterial genetics or a teacher looking for a detailed explanation to share with your class, this article will provide a thorough understanding of what is an operon, its structure, function, and significance in gene regulation. Let’s explore the operon definition, its types, and how it controls gene expression in prokaryotic cells.

Table of Contents
Operon Definition
An operon is a functional unit of DNA in prokaryotic organisms that consists of a cluster of structural genes controlled by a single promoter region. These genes are transcribed together as a single mRNA molecule, allowing for coordinated expression. The operon model was first introduced by François Jacob and Jacques Monod in 1961, based on their studies of the lac operon in E. coli.
In simpler terms, what is an operon? It is a set of genes that work together under the control of a single regulatory system. This system includes a promoter region, an operator site, and structural genes. The operon function is to ensure that genes involved in related processes are expressed simultaneously, optimizing the cell’s response to environmental changes.
Operon Meaning and Structure
The term operon refers to a set of functionally related genes that are transcribed as a single messenger RNA (mRNA). This system enables bacteria to adapt to environmental changes by coordinating gene expression efficiently.
Components of an Operon
An operon structure consists of the following elements:
| Component | Function |
|---|---|
| Promoter Region | This is the site where RNA polymerase binds to initiate transcription. |
| Operator Site | The regulatory sequence is where repressor or activator proteins bind to control gene expression. |
| Structural Genes | These genes encode proteins necessary for a specific metabolic pathway. |
| Regulatory Genes | Encode proteins (such as repressors) that regulate the operon’s activity. |
These components work together to ensure efficient transcriptional regulation in bacteria.
What is the purpose of op-erons in gene expression?
The primary function of operons in bacterial gene regulation is to control gene expression, ensuring proteins are synthesized only when needed. This mechanism provides several advantages:
Why is Operon Regulation Important?
- Energy Conservation: Prevents wasteful protein synthesis.
- Rapid Environmental Response: Allows bacteria to adapt to changing nutrient availability.
- Efficient Gene Expression: Coordinates multiple genes involved in the same biological process.
For example, the lac operon activates only when lactose is present, ensuring enzymes for lactose metabolism are produced only when required. Similarly, the trp operon controls tryptophan biosynthesis based on amino acid availability.
Operon Function: How Operons Control Gene Expression
The primary function of operons is to regulate gene expression in prokaryotic cells efficiently. By grouping related genes together, operons allow for coordinated transcription and translation, ensuring that the cell produces the necessary proteins only when needed. This mechanism is crucial for conserving energy and resources.
How Operons Work
- Transcriptional Control: The operon is activated or repressed based on the presence or absence of specific molecules.
- Regulatory Proteins: There are two kinds of proteins that bind to the operator site and control how RNA polymerase can get to the structural genes. These are called repressors and activators.
- Environmental Signals: Operons respond to environmental changes, such as nutrient availability, by turning genes on or off.
For example, the lac operon in E. coli is activated in the presence of lactose, allowing the bacteria to metabolize the sugar. On the other hand, when tryptophan is abundant, the trp operon is turned off, which stops the amino acid from being made when it is not needed.
Types of Operons in Bacterial Gene Regulation
Operons can be classified into two main types based on their regulation:
- Inducible Operon: These operons are usually off but can be turned on in the presence of a specific inducer molecule. The lac operon is a classic example of an inducible operon.
- Repressible Operon: These operons are typically on but can be turned off when a specific molecule is abundant. The trp operon is an example of a repressible operon.
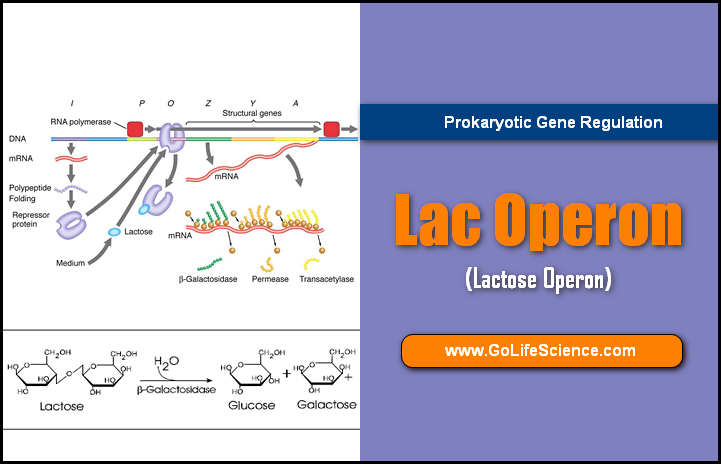
Lac Operon (Inducible Operon)
The lac operon in E. coli controls the metabolism of lactose and is a classic example of an inducible operon.
| Component | Function |
|---|---|
| LacZ | Encodes beta-galactosidase, which breaks down lactose into glucose and galactose. |
| LacY | Encodes lactose permease, which facilitates lactose transport into the cell. |
| LacA | Encodes thiogalactoside transacetylase, involved in detoxification. |
| LacI (Repressor) | Binds to the operator site, blocking transcription when lactose is absent. |
When lactose is present, it binds to the LacI repressor, preventing it from binding to the operator. This allows RNA polymerase to transcribe the lac genes, leading to enzyme production for lactose metabolism.
Trp Operon (Repressible Operon)
The trp operon regulates tryptophan biosynthesis in E. coli and functions as a repressible operon.
| Condition | Effect |
| High tryptophan | The repressor binds to the operator, blocking transcription. |
| Low tryptophan | The repressor is inactive, allowing transcription to proceed. |
When tryptophan levels are high, it binds to the trp repressor, enabling it to attach to the operator and inhibit transcription. When tryptophan is scarce, the repressor remains inactive, allowing gene expression for tryptophan synthesis.
Role of Operons in Bacterial Gene Regulation
The role of operon in gene regulation is to ensure that genes are expressed only when needed. This regulation occurs at the transcriptional level, where regulatory proteins interact with the operator site to control the binding of RNA polymerase to the promoter region.
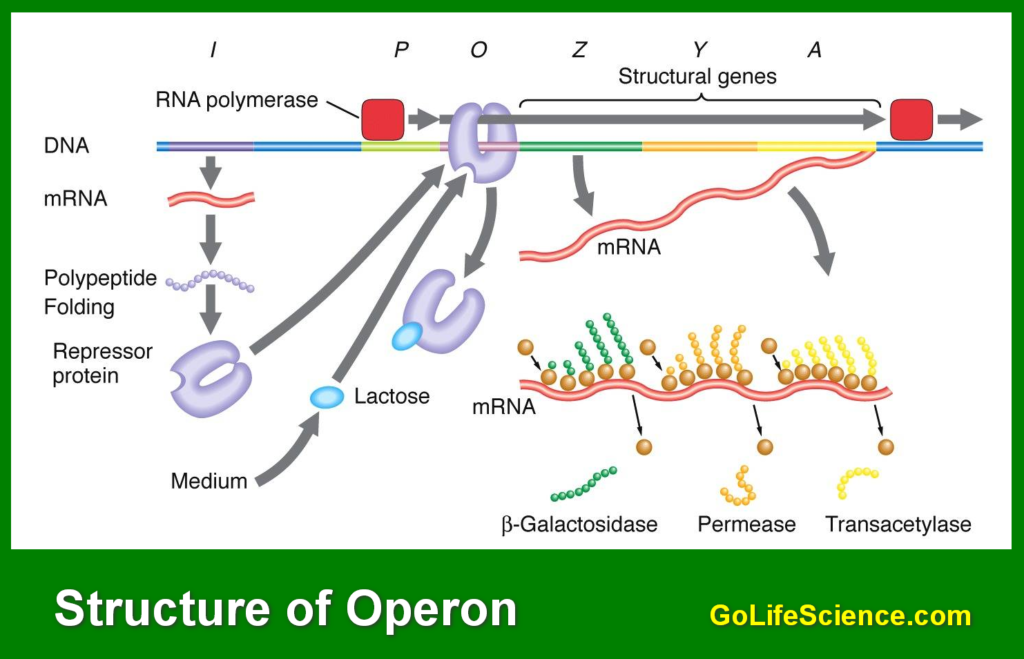
Operons play a crucial role in gene regulation, functioning as molecular switches that control transcription.
| Term | Definition |
| Operon Activation | When an operon allows transcription to occur. |
| Operon Repression | When an operon blocks transcription through a repressor protein. |
| Regulatory Proteins | Proteins that influence operon activity. |
| Gene Clusters | Groups of genes controlled by a single promoter. |
Key Mechanisms
- Operon Activation: In the absence of a repressor, RNA polymerase binds to the promoter, initiating transcription.
- Operon Repression: When a repressor binds to the operator, RNA polymerase is blocked, preventing transcription.
This mechanism allows prokaryotic cells to adapt quickly to changing environments, optimizing their metabolic processes.
Examples of Operons: Lac and Trp Operons
1. Lac Operon
The lac operon is a well-studied example of an inducible operon. It consists of three structural genes (lacZ, lacY, and lacA) involved in lactose metabolism. When lactose is present, it binds to the repressor protein, causing it to release from the operator. This allows RNA polymerase to transcribe the genes, enabling the cell to metabolize lactose.
2. Trp Operon
The trp operon is an example of a repressible operon. It contains genes responsible for synthesizing tryptophan. When tryptophan levels are high, it acts as a corepressor, binding to the repressor protein and enabling it to block transcription. This prevents the unnecessary production of tryptophan.
Operon Biology Definition: Why Are Operons Important?
The operon biology definition highlights its significance in prokaryotic gene expression. Operons are crucial for:
- Efficient Gene Regulation: They allow cells to respond quickly to environmental changes.
- Energy Conservation: By producing proteins only when needed, cells save energy.
- Simplified Genetic Control: Operons provide a streamlined mechanism for controlling multiple genes simultaneously.
Operon Meaning in Genetic Control Mechanisms
The operon meaning extends beyond simple gene clusters. It represents a sophisticated genetic control mechanism that ensures the precise regulation of metabolic pathways. By understanding how operons control gene expression, scientists can manipulate these systems for applications in biotechnology, medicine, and agriculture.
Operon Model: A Framework for Understanding Gene Regulation
The operon model provides a framework for understanding how genes are regulated in prokaryotes. It emphasizes the importance of regulatory proteins, promoter regions, and operator sites in controlling transcriptional control. This model has been instrumental in advancing our knowledge of bacterial genetics and gene clusters.
Conclusion
In summary, what is the purpose of operons in protein synthesis? Operons streamline the process of gene regulation, ensuring that proteins are synthesized only when required. By grouping related genes together and controlling their expression through regulatory proteins, operons enable prokaryotic cells to adapt efficiently to their environment.
Whether you’re exploring the lac operon, the trp operon, or the broader concept of operon structure, understanding operons is essential for grasping the fundamentals of prokaryotic gene expression and genetic control mechanisms.
Understanding what is an operon in biology is essential for comprehending bacterial gene regulation. The operon function allows bacteria to regulate gene expression efficiently, ensuring they produce proteins only when needed. By studying the operon model, scientists have gained valuable insights into transcriptional control and gene regulation mechanisms, which are fundamental in molecular biology, genetics, and biotechnology
Frequently Asked Questions (FAQs)
What is an operon in biology?
An operon is a cluster of genes in prokaryotic cells that are transcribed together and regulated by a single promoter and operator.
What is the operon function?
The operon’s job is to control gene expression by making sure that related genes are transcribed together in response to signals from the outside world.
What are the types of operons?
The two main types are inducible operons (e.g., lac operon) and repressible operons (e.g., trp operon).
How do operons control gene expression?
Operons control gene expression by interacting with regulatory proteins that bind to the operator site and either let RNA polymerase copy the structural genes or stop it.
What is the role of operon in gene regulation?
Operons play a crucial role in gene regulation by ensuring that genes are expressed only when needed, optimizing the cell’s response to environmental changes.