Enzymes are the biological catalysts that drive almost every chemical reaction in living cells. While many enzymes simply speed up reactions, some have a special ability to act like metabolic regulators. These are called allosteric enzymes.
Unlike ordinary enzymes, allosteric enzymes possess an extra regulatory site (allosteric site) along with their active site. This unique feature allows them to respond to specific effectors—molecules that either activate or inhibit their function.
These enzymes undergo a conformational change to rapidly switch between active and inactive states. This ability makes them vital for maintaining metabolic balance.
Allosteric enzymes are especially important at rate-limiting steps of metabolic pathways such as glycolysis, nucleotide biosynthesis, and nitrogen metabolism. They guarantee that a pathway does not “overwork” when energy or biomolecules are already in surplus.
Beyond basic biology, understanding allosteric enzymes has practical value in medicine and drug design. Many modern drugs target the allosteric sites of enzymes. This approach helps in fine-tuning activity. It provides greater specificity and fewer side effects compared to traditional inhibitors.

What are Allosteric Enzymes?
Allosteric enzymes are a special group of enzymes. They regulate the rate of metabolic pathways by binding molecules at a site other than the active site. This additional site, called the allosteric site (or regulatory site), controls the enzyme’s activity by changing its shape and function.
In simple terms, ordinary enzymes follow classic Michaelis–Menten kinetics. They show a smooth hyperbolic activity curve. In contrast, allosteric enzymes display sigmoidal kinetics (S-shaped curve).
This occurs because they undergo cooperative binding. This means the binding of one substrate or effector makes it easier (or harder) for the next one to bind.
These enzymes often serve as control points in biochemical pathways. They ensure that processes like glycolysis, nucleotide biosynthesis, and nitrogen metabolism run efficiently. This prevents the waste of energy or resources.
Difference Between Allosteric and Regular Enzymes
| Feature | Regular (Non-Allosteric) Enzymes | Allosteric Enzymes |
|---|---|---|
| Regulation | Activity controlled only at active site | Controlled by both active site and allosteric site |
| Kinetics Curve | Hyperbolic (Michaelis–Menten type) | Sigmoidal (S-shaped, due to cooperativity) |
| Structure | Usually single-subunit proteins | Typically multisubunit (quaternary structure) |
| Binding | Independent substrate binding | Cooperative substrate/effector binding |
| Role in Metabolism | Catalysis only | Act as regulators of metabolic pathways |
| Examples | Hexokinase, Catalase | PFK-1, ATCase, Pyruvate kinase |
Features of Allosteric Enzymes
- Multisubunit nature: Most allosteric enzymes consist of several polypeptide chains, allowing cooperative interactions.
- Allosteric site present: In addition to the catalytic active site, they have a regulatory site for effectors.
- Positive and negative effectors:
- Positive effectors (activators): Increase enzyme activity (e.g., AMP, ADP).
- Negative effectors (inhibitors): Decrease enzyme activity (e.g., ATP, citrate).
- Sigmoidal kinetics: Enzyme activity shows an S-shaped curve due to cooperativity.
- Switch-like behavior: Small changes in effector concentration can rapidly shift the enzyme between active (R state) and inactive (T state).
- Regulatory role: Usually found at rate-limiting steps of pathways, controlling the flow of metabolites.
Structural & Functional Features of Allosteric Enzymes
The remarkable behavior of allosteric enzymes is rooted in their unique structural features. Unlike simple enzymes, they often have multiple subunits.
They have special binding sites that allow them to sense and respond to metabolic signals. Let’s break down the key structural elements that explain their regulatory power.
a. Active Site vs Allosteric Site
Before diving into regulation, it’s essential to understand that allosteric enzymes contain two distinct functional sites.
While the active site performs the core catalytic reaction, the allosteric site acts like a control switch. Together, these sites ensure that enzyme activity is tuned according to the cell’s needs.
- Active site → Region where the substrate binds and the chemical reaction occurs.
- Allosteric site (regulatory site) → A separate pocket where allosteric effectors bind.
- Positive effectors (activators): Stabilize the enzyme in its active form.
- Negative effectors (inhibitors): Shift the enzyme into its inactive form.
📌 Important: This two-site system explains why allosteric regulation is more dynamic than simple substrate binding.
b. Multisubunit Structure
Most allosteric enzymes are oligomeric proteins composed of multiple subunits. This arrangement is not just for stability—it is the foundation of cooperativity and regulation.
- Each subunit have an active site.
- Binding in one subunit influences the shape of others (subunit interaction).
- Provides the quaternary structure flexibility needed for allosteric transitions.
Example: Aspartate transcarbamoylase (ATCase) has 12 subunits—6 catalytic and 6 regulatory—making it a classic model of allosteric behavior.
c. Tense (T) vs Relaxed (R) States
To understand how allosteric regulation works, it’s important to know that enzymes do not exist in a fixed shape. Instead, they can flip between two main conformations: a low-affinity inactive form and a high-affinity active form. These are called the Tense (T) state and the Relaxed (R) state.
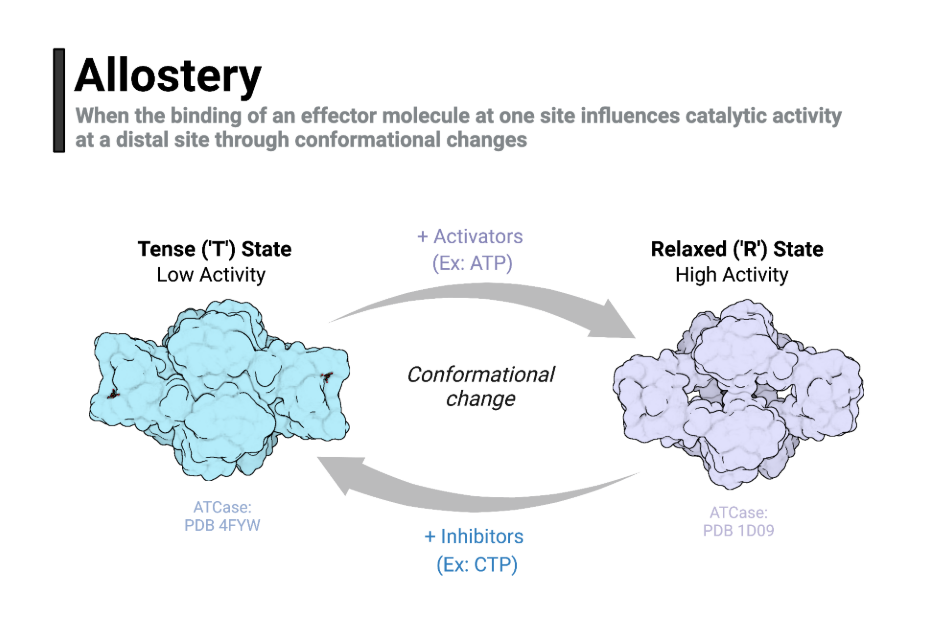
- Tense (T) state
- Low affinity for substrate.
- Enzyme remains mostly inactive.
- Stabilized by inhibitors like ATP or CTP.
- Relaxed (R) state
- High affinity for substrate.
- Enzyme becomes fully active.
- Stabilized by activators like AMP or ADP.
This T ↔ R transition is the molecular basis of allosteric regulation. It allows enzymes to respond quickly to changes in the cellular environment.
Quick Revision Table – Structural & Functional Features
| Feature | Explanation | Key Point |
|---|---|---|
| Active vs Allosteric Site | Active site → catalysis; Allosteric site → regulation by effectors | Dual-site system |
| Multisubunit Structure | Made of several polypeptide chains forming quaternary structure | Basis for cooperativity |
| Cooperativity | Binding at one subunit influences binding at others | Explains sigmoidal kinetics |
| Tense (T) State | Low substrate affinity; stabilized by inhibitors | Inactive form |
| Relaxed (R) State | High substrate affinity; stabilized by activators | Active form |
Mechanisms of Allosteric Regulation
Allosteric enzymes do more than switch on or off. They follow precise molecular models. These models explain how binding at one site influences the entire protein.
These models help us understand phenomena like cooperativity, feedback inhibition, and enzyme activation. Let’s explore the three most important models of allosteric regulation.
a. Concerted Model (MWC Model)
The concerted model, also known as the Monod-Wyman-Changeux (MWC) model, describes how enzyme subunits transition. All subunits switch between the T (tense) state and the R (relaxed) state together.
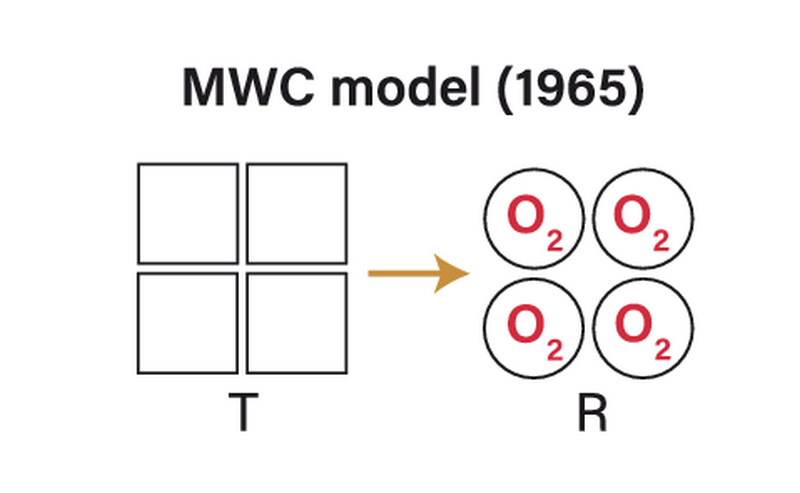
- All subunits are in the same state at any given time.
- Binding of one substrate molecule increases the chance of all subunits flipping to the R state.
- Explains positive cooperativity seen in enzymes like aspartate transcarbamoylase (ATCase).
- Used to describe classic sigmoidal kinetics in allosteric enzymes.
b. Sequential Model (KNF Model)
The sequential model, also called the Koshland-Némethy-Filmer (KNF) model, explains that subunits do not all switch states together. Instead, they switch one at a time.
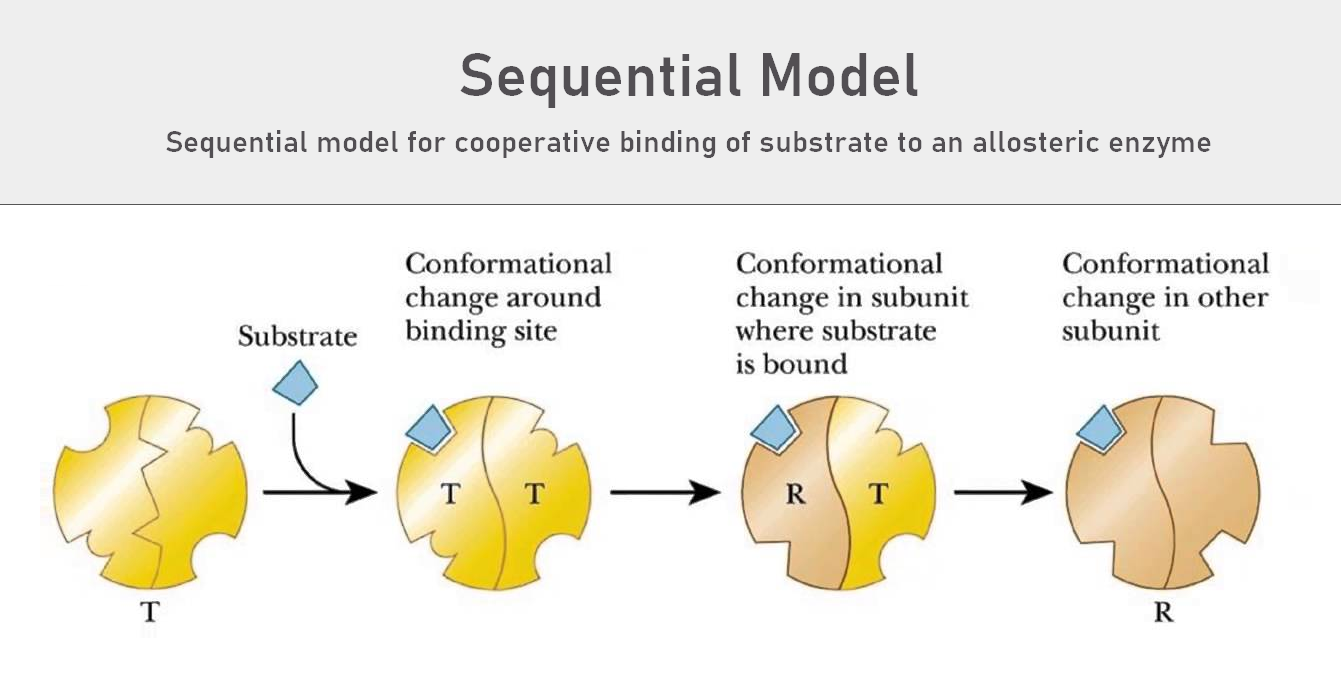
- Substrate binding to one subunit induces a conformational change.
- This change slightly increases the affinity of neighboring subunits.
- Allows for a stepwise transition between T and R states.
- Better explains negative cooperativity (binding reduces affinity at other sites).
- Common in enzymes that display variable cooperative behavior.
c. Ensemble Model of Allostery (Modern)
A more modern view of allosteric regulation is the ensemble model. This model considers enzymes as dynamic molecules. They have multiple conformations existing at the same time.

- Instead of fixed states, enzymes exist as a population of conformations.
- Substrate or effector binding shifts the population balance toward active or inactive forms.
- Explains complex regulatory behaviors beyond simple MWC or KNF models.
- Provides insights into drug design targeting allosteric enzymes in medical research.
Together, these models highlight that allosteric regulation is flexible, adaptive, and essential for fine-tuned metabolic control.
Examples of Allosteric Enzymes in Metabolism
The real beauty of allosteric enzymes is seen in metabolism, where they act as control points to maintain cellular balance.
By responding to activators and inhibitors, these enzymes ensure that energy production, biosynthesis, and nutrient utilization are optimized. Let’s look at some well-studied examples.
1. Phosphofructokinase-1 (PFK-1)
PFK-1 is one of the most famous allosteric enzymes in glycolysis, regulating the conversion of fructose-6-phosphate to fructose-1,6-bisphosphate. It serves as a metabolic gatekeeper for glucose breakdown.
- Pathway: Glycolysis
- Activators: AMP, ADP, fructose-2,6-bisphosphate
- Inhibitors: ATP, citrate
- Role: Acts as an energy sensor, accelerating glycolysis when energy is low and slowing it when energy is abundant.
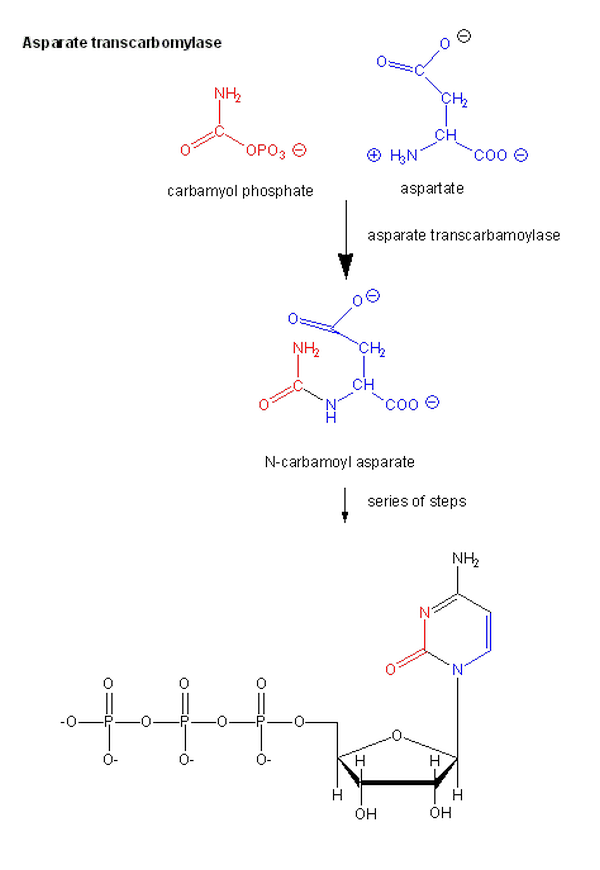
2. Aspartate Transcarbamoylase (ATCase)
ATCase controls the first step in pyrimidine biosynthesis. It balances nucleotide production, which is critical for DNA and RNA synthesis.
- Pathway: Pyrimidine biosynthesis
- Activators: ATP
- Inhibitors: CTP (end-product inhibition)
- Role: Maintains nucleotide balance between purines and pyrimidines, preventing wasteful overproduction.
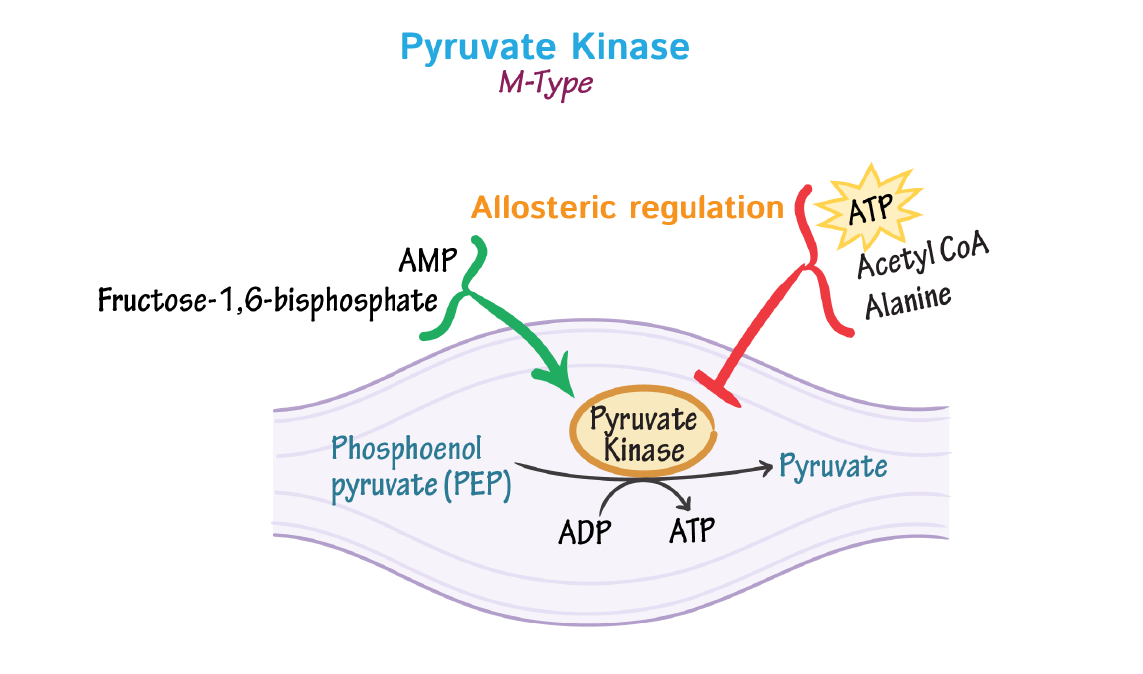
3. Pyruvate Kinase
Pyruvate kinase catalyzes the final step of glycolysis, converting phosphoenolpyruvate (PEP) to pyruvate. Its regulation ensures that glycolysis moves smoothly toward energy production.
- Pathway: Glycolysis
- Activators: Fructose-1,6-bisphosphate (feed-forward activator)
- Inhibitors: ATP, alanine
- Role: Coordinates glycolytic flux, preventing unnecessary pyruvate buildup when energy is sufficient.
4. Glutamine Synthetase
Glutamine synthetase plays a central role in nitrogen metabolism, synthesizing glutamine from glutamate and ammonia. It is heavily regulated by multiple feedback inhibitors.
- Pathway: Nitrogen metabolism
- Activators: None (primarily feedback inhibited)
- Inhibitors: End-products like glycine, alanine, AMP, carbamoyl phosphate
- Role: Acts as a metabolic hub, adjusting nitrogen assimilation according to cellular needs.
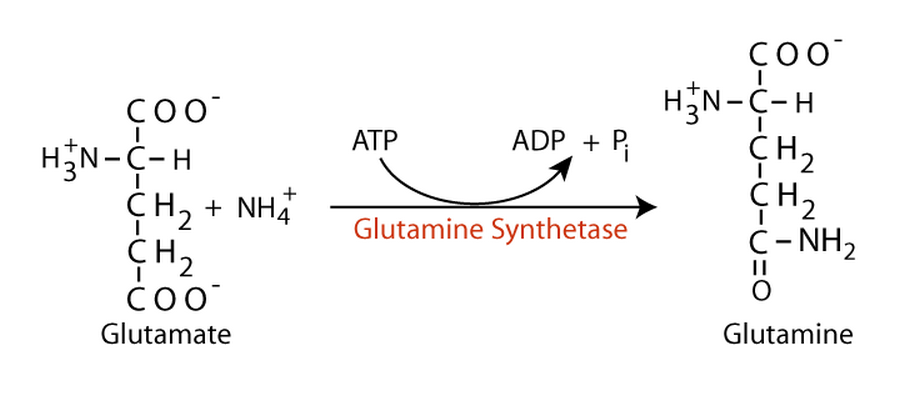
Table: Key Allosteric Enzymes, Regulators, and Functions
| Enzyme | Pathway | Activators | Inhibitors | Role in Metabolism |
|---|---|---|---|---|
| PFK-1 | Glycolysis | AMP, ADP, Fructose-2,6-BP | ATP, Citrate | Energy sensor controlling glycolytic flux |
| ATCase | Pyrimidine biosynthesis | ATP | CTP | Balances nucleotide synthesis |
| Pyruvate Kinase | Glycolysis | Fructose-1,6-Bisphosphate | ATP, Alanine | Regulates final step of glycolysis |
| Glutamine Synthetase | Nitrogen metabolism | — | Glycine, Alanine, AMP, Carbamoyl phosphate | Maintains nitrogen balance and amino acid pools |
These examples highlight how allosteric regulation is central to metabolic control, linking energy status, biosynthetic needs, and cellular growth.
Clinical and Drug Design Relevance
Allosteric enzymes are not only fascinating in the classroom—they are also game changers in medicine and biotechnology.
Their unique property of having separate regulatory sites makes them highly attractive for drug discovery and therapeutic interventions. Unlike traditional drugs that target the active site, allosteric modulators offer greater selectivity.
They result in fewer side effects and have the ability to fine-tune enzyme activity rather than completely shutting it down.
Why Allosteric Enzymes Are Important in Drug Discovery
- Specificity: Allosteric modulators bind to unique sites, reducing off-target interactions.
- Tunable activity: They can enhance or inhibit enzyme function without fully blocking the enzyme.
- Reduced resistance: Drugs targeting allosteric sites often avoid the resistance seen with traditional inhibitors.
- Novel therapeutic pathways: Many diseases are linked to dysregulated enzyme activity, and allosteric regulation provides new solutions.
Case Studies in Therapeutics
a. Allosteric Modulators in Diabetes
- Glucokinase (GK) activators are explored for type 2 diabetes treatment.
- GK acts as a glucose sensor in the pancreas and liver.
- GK activators (GKAs) enhance insulin secretion and glucose utilization.
- PFK-1 modulation is also under study for controlling glycolysis in diabetic conditions.
b. Cancer Therapy Targeting Allosteric Enzymes
- Cancer cells rely heavily on altered metabolism (Warburg effect).
- Targeting isocitrate dehydrogenase (IDH1/IDH2) with allosteric inhibitors blocks abnormal metabolic flux.
- MEK1/2 allosteric inhibitors are already in clinical use for certain cancers (e.g., melanoma).
- This approach reduces tumor growth while sparing normal cells.
c. Antibiotics Targeting Bacterial Allosteric Enzymes
- Some antibiotics disrupt bacterial metabolism by targeting allosteric sites.
- Example: DHPS inhibitors block folate synthesis in bacteria.
- Allosteric enzyme modulation provides a strategy to overcome antibiotic resistance.
Recent Research Updates 2025
- Allosteric drugs are gaining attention in precision medicine (source: Nature Reviews Drug Discovery, 2024).
- Over 20 FDA-approved allosteric modulators are now available for diseases ranging from epilepsy to cancer.
- Ongoing research explores AI-driven drug design to predict new allosteric sites.
- Biotechnology companies are actively patenting allosteric modulators for metabolic and neurological disorders.
In summary, allosteric enzymes are powerful drug targets, offering flexibility, precision, and innovation in modern therapeutics.
Applications & Importance in Biochemistry
Allosteric enzymes are not just classroom concepts—they are vital players in real biological systems. By acting as metabolic regulators and molecular switches, they ensure smooth control of biochemical pathways.
Their unique properties also make them highly valuable in biotechnology, medicine, and drug design. Let’s explore their key applications.
1. Regulation of Metabolism
- Act as rate-limiting enzymes in critical pathways like glycolysis and nucleotide biosynthesis.
- Respond to cellular energy status (ATP/AMP balance).
- Prevent unnecessary energy loss by feedback inhibition.
- Example: PFK-1 regulating glycolysis.
2. Signal Transduction Pathways
- Participate in cell signaling cascades, ensuring proper communication inside the cell.
- Enable rapid responses to environmental or hormonal changes.
- Example: Enzymes involved in G-protein signaling pathways.
3. Synthetic Biology & Biotechnology
- Engineered as biosensors to detect small molecules.
- Used to build synthetic regulatory circuits in metabolic engineering.
- Help optimize biofuel and bioproduct synthesis by controlling pathway flux.
4. Pharmaceutical Applications
- Allosteric modulators are powerful tools in drug design.
- Provide higher specificity than active-site inhibitors.
- Reduce chances of resistance in diseases like cancer and bacterial infections.
- Examples: MEK inhibitors in cancer therapy, glucokinase activators in diabetes.
Table: Applications of Allosteric Enzymes
| Application Area | Key Role / Example | Importance |
|---|---|---|
| Metabolic Regulation | PFK-1 in glycolysis, ATCase in nucleotide biosynthesis | Maintains energy balance and metabolic control |
| Signal Transduction | G-protein–linked enzyme regulation | Ensures accurate cellular communication |
| Biotechnology | Engineered allosteric enzymes as biosensors | Applied in synthetic biology and bioengineering |
| Pharmaceuticals | MEK inhibitors, Glucokinase activators | Drug design with high specificity, fewer side effects |
In short, allosteric enzymes are master regulators—they guide metabolism, signal processing, and even inspire new medical and biotechnological innovations.
Frequently Asked Questions (FAQs) on Allosteric Enzymes
What are allosteric enzymes in simple terms?
Allosteric enzymes are special proteins that help regulate chemical reactions in cells. Unlike normal enzymes, they have two sites. One is an active site for the substrate. The other is an allosteric site for molecules that can either activate or inhibit the enzyme. This allows the enzyme to act like a switch, turning activity up or down as needed. An example is PFK-1, which regulates glycolysis.
How do allosteric enzymes differ from normal enzymes?
Normal enzymes follow simple Michaelis-Menten kinetics and usually respond only to substrate concentration. Allosteric enzymes, on the other hand, show sigmoidal kinetics. The binding of a molecule to one subunit affects the binding on others. This phenomenon is called cooperativity. They are often multisubunit proteins. They can be regulated by activators and inhibitors at the allosteric site. This regulation gives them a greater role in controlling metabolic pathways.
Why are allosteric enzymes important in metabolism?
Allosteric enzymes act as key regulators of metabolic pathways. They are often found at rate-limiting steps, ensuring that the cell does not produce excess metabolites. They respond to changes in energy levels, such as the ratio of ATP to AMP. In this way, they help balance energy production. They also manage resource use. For example, PFK-1 in glycolysis speeds up or slows down glucose breakdown depending on the cell’s energy needs.
What is the difference between an activator and an inhibitor in allosteric regulation?
An allosteric activator is a molecule. It binds to the regulatory site and increases the enzyme’s activity. This is often achieved by stabilizing its relaxed (R) state. In contrast, an allosteric inhibitor binds to the same regulatory site. It decreases enzyme activity by favoring the tense (T) state. For instance, AMP activates PFK-1 to speed up glycolysis, while ATP inhibits it when energy is abundant.
How are allosteric enzymes used in drug design?
Allosteric enzymes are ideal targets for drugs. Molecules can bind to the regulatory site rather than the active site. This allows drugs to fine-tune enzyme activity with high specificity and fewer side effects. Many modern therapies exploit allosteric regulation. Cancer treatments like MEK inhibitors use this approach. Diabetes drugs like glucokinase activators also use it to control metabolic and signaling pathways efficiently.
Conclusion
In summary, allosteric enzymes are unique regulatory proteins. They play a central role in metabolic pathways. They do this by controlling the flow of biochemical reactions. Their special allosteric site allows them to respond to activators and inhibitors. This response causes conformational changes.
These changes switch the enzyme between the Tense (T) and Relaxed (R) states. This makes them crucial for energy management, biosynthesis regulation, and maintaining cellular homeostasis.
Understanding the mechanisms of allosteric regulation is important. This includes the MWC (concerted) model, KNF (sequential) model, and ensemble (population-shift) model. These mechanisms help students appreciate how enzymes act as biological switches.
Key examples include PFK-1, ATCase, pyruvate kinase, and glutamine synthetase. They show the practical relevance of allosteric enzymes in glycolysis, nucleotide biosynthesis, and nitrogen metabolism.
From a modern perspective, allosteric enzymes are also vital in drug design, disease treatment, and biotechnology. They offer highly specific targets for therapeutic intervention. These enzymes also allow innovative biotechnological applications.
For students, these points serve as easy-to-revise notes that simplify complex concepts like cooperative binding, sigmoidal kinetics, and feedback regulation.